
I enjoy writing (or, at least, the idea of writing) about a veritable plethora of topics: philosophy, religion, politics, economics, society. However, my education is actually in the sciences – I am a published biochemist. This has made me strive for scientific accuracy, or at least scientific plausibility, in my fiction writing. For science fiction, the CRISPR/Cas9 system of gene editing leads to all sorts of possibilities, some of which I explore in my Incarnate novel series (shameless plug – book 2 coming soon!). In my novel, people use CRISPR/Cas9 to modify their somatic cells in order to give themselves practical biochemical upgrades (ie having one’s fingerprint pattern shift slowly over time so that the person cannot be tracked) to more aesthetic ones (ie giving oneself a series of subdermal bioluminescent vessels that can be lit up at will).
But if one wants to know how to better understand the possibilities and limitations to CRISPR/Cas9 (at least in its current state in August of 2018 – my novel series takes place in the future where some of the practical limitations will hopefully be overcome), it is necessary to understand how it actually works. Here, I am going to lay out in hopefully simple terms (although this may be somewhat technical at times) what CRISPR/Cas9 is, how it works, and what people are currently using it for (or debating whether they ought to use it for these things, anyway). This will not be an exhaustive review of everything CRISPR/Cas9 related, but it will hopefully make it understandable enough for the interested laity to know what it is and for fellow sci-fi writers to use it more effectively in their writing.
A Few Basic Preliminary Genetics Concepts (skippable)
DNA, which stands for deoxyribonucleic acid, is a long polymer composed of nucleotides containing one of four different nitrogenous bases called adenine (A), guanine (G), thymine (T) (uracil (U) in RNA), and cytosine (C). Nucleotides are composed of the nitrogenous bases listed above bound to a ribose sugar (deoxyribose in DNA, ribose in RNA) bound to a phosphate.

Two single stranded DNA polymers then form into a double stranded DNA duplex, which generates the double helical structure famously discovered by Watson and Crick in 1953 (using Rosalind Franklin’s x-ray crystal structures). We know from Chargaff’s Rules that adenine (A) always binds to thymine (T) and that guanine (G) always binds to cytosine (C). This means that the two strands must have complementary sequences running in anti-parallel directions, as depicted below.

What this also means is that anything that will bind to the blue sequence in the figure above must have the same sequence as that shown in red. Sequences close to it, for example a single base pair off, will still be able to bind, but with less affinity. The further away the sequences get from 100% complementarity, the less they will bind and the more they will unbind.
The bases in each strand, such as that between the first adenine (A) and cytosine (C) at the 3` end of the blue strand, are held together by a phosphodiester bond when the 3` oxygen binds to the phosphate group on the next base, circled in the figure below.

These phosphodiester bonds can be broken using enzymes. If the enzyme has to start from the end, such as breaking the A off from the C at the 3`of the blue strand, the enzyme is classified as an exonuclease (the suffix -ase in biochemistry always denotes an enzyme) and is said to have exonuclease activity. If it can break bonds internally, such as between the T and G in the third and fourth positions in the blue strand, the enzyme is classified as an endonuclease. and is said to have endonuclease activity.

What did CRISPR/Cas9 Originally Evolve to do?
CRISPR, which stands for Clustered Regularly Interspaced Short Palindromic Repeats, evolved as a natural immune system that prokaryotes (bacteria) use to fight phage (virus) infection. Phages work by injecting their own DNA into a bacteria and then hijacking the cell’s replication system.

Barrangou et al discovered in 2007 that some bacteria (in their case, the Streptococcus thermophilus used to make yogurt) can incorporate segments of viral DNA into their own genome if they are exposed to the virus. This is done by the Cas1-Cas2 protein complex (Cas stands for CRISPR associated since they are CRISPR associated proteins) whereby the Cas1 acts as an endonuclease (it cuts nucleic acid strands, in this case deoxyribonucleic acid, or DNA) to cut out small sections of the viral DNA (adjacent to a PAM site – more on this later). This process generates what are known as spacers (a small section of the viral DNA sequence). These spacers are then inserted into the CRISPR locus of the bacterial genome (specific site in the DNA genome of the bacteria). Multiple spacers can be stored in this locus, which grants the bacteria a defense against a variety of viral strains.
The bacteria can then transcribe this small section of DNA into RNA segments. These segments will be the complement sequence to the original viral DNA sequence, which means that they can bind with the viral DNA. Bacteria use these RNA fragments to guide the Cas9 protein toward the specific sequence in viral DNA upon infection by the same viral strain (thus, these RNA fragments are known as the guide RNA or gRNA). The Cas9 can then cut the viral DNA, rendering it inert until other nucleases can degrade it, thereby halting the infection.

How is the CRISPR/Cas9 System Able to be Exploited for Genome Editing?
Since 1971 the restriction enzyme system, which cuts DNA at specific palindromic sequences (and, interestingly, also used by bacteria as a defense against viral infection) has been the tool for molecular cloning (moving sequences of DNA from one genome to another, within or between species). This method has limitations because it requires the specific palindromic sequence in the DNA for it to make its cut, such as the AATT sequence for EcoRI (see figure below). These palindromic sequences A) are not always present in the genome and B) may be present in places you do not want to cut (this latter part I have found particularly annoying in my own research). Thus, the multiple cloning site is popular for molecular cloning, which is a prerequisite to just about anything one does in biochemistry.

With CRISPR/Cas9, though, the use of the guide RNA can bring the Cas9 protein to precise, specific sites in the genome, no palindrome required (Gasiunas et al, 2012) (Jinek et al, 2012). Using synthetic guide RNAs with a specific desired sequence could then be used to cut the genome wherever you wanted (suggested by Jinek et al, 2012 and demonstrated by Cong et al, 2013).
What are the Molecular Mechanisms of CRISPR/Cas9? (technical)
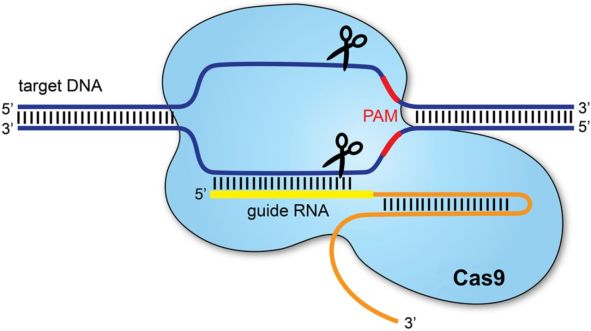
The gRNA is actually composed of two different RNA strands that are necessary for Cas9 association with the cut site, called CRISPR RNA (crRNA) and trans-activating CRISPR RNA (tracrRNA). Both are needed in order for Cas9 to bind to the target sequence. The crRNA contains the 20 base pair guide sequence that is complementary to one strand of the DNA, the target sequence, and the tracrRNA orients the crRNA in order to properly associate with the complementary strand.

The Cas9-crRNA-tracrRNA complex scans the genome for the Protospacer Adjacent Motif (PAM) sequence, whereby arginine residues in the PAM Interacting (PI) domain of the Cas9 protein will interact with the guanine (G) bases in the PAM sequence and a lysine in the PI domain will bind in the DNA duplex minor groove. This will allow the guide sequence of the crRNA to interact with the DNA duplex, and if the crRNA sequence matches with the complementary strand, it will bind to it.
Next, the Cas9 protein uses its HNH endonuclease domain to cleave the complementary strand 3 bases upstream of the PAM site. The Cas9 H840 residue deprotonates a water molecule, which then attacks the phosphate in the phosphodiester bond, which is made more electrophilic by coordination with a magnesium ion. The triganol bipyramidal transition state then collapses back down into the energetically more favorable tetrahedral geometry by regenerating a double bond with an oxygen and kicking off the 3′ – 5′ phosphate bond, separating the two adjacent bases.
A similar RuvC endonuclease activity is used to cleave the noncomplementary strand 3 to 8 bases upstream of the PAM site, using the H983 residue to activate the water molecule.

Above is a generic illustration of phosphodiester bond cleavage. In the specific cases of the HNH and RuvC endonuclease activities, X = the Histidine (H) residue used to activate the water molecule; Z = a different water molecule (H2O) from solvent; and then R = the rest of the DNA sequence on either side of the phosphodiester bond in question.
What’s with that PAM Sequence?
So, what about this pesky PAM sequence that keeps coming up that I promised to talk about earlier? The Protospacer Recognition Motif (PAM) is a three base pair sequence that must be near the target site in the genome. There is variation in this sequence based on bacterial strain. The Cas9 protein binds to this sequence as it scans the genome, stopping in order for the guide RNA to attempt to bind to the DNA. If the DNA has no sequence complementarity with the gRNA, the complex will simply move on. If the DNA sequence does complement the gRNA guide sequence, then Cas9 will stop long enough to cleave both strands at the cut site.
The PAM sequence is a limitation for gene editing in that any genome editing must be adjacent to this sequence. Using Cas9 from different species, or altering the PAM specificity of Cas9 using guided evolution, are being explored as ways to get around this limitation.
Genome Editing with CRISPR/Cas9
Now that we have cut our DNA strand, what do we do next? Now we can insert or remove precisely targeted DNA from a complete genome. This can be done using homology directed repair to insert new genetic material into the part of the genome that has been cut.

Then comes the fun part.
What Can CRISPR/Cas9 be Used For?
The CRISPR/Cas9 system can be used for all sorts of real world and sci-fi things. Here is just a few that I can come up with.
Knockdowns for scientific research
A huge part of genetic research is finding out what a specific gene does. This can’t really be done by just looking at the protein it codes for in isolation, it has to be studied in context. The method often used is called a knockdown. This is done by greatly decreasing or completely eliminating the expression of the gene in a cellular or organism context. The researcher then either observes the cell or organism phenotype compared to wild type (cell or organism that has not had that gene’s activity decreased or eliminated) or performs biochemical assays to measure things like mRNA transcription (eg groSEQ), mRNA concentrations at time points after treatment (eg qPCR), differential protein concentrations (eg immunoprecipitation pull-down followed by Western blot), and so on between treated and wild type.
Science fiction idea: wouldn’t a toxin that knocks down genes be a particularly cruel way to kill someone? The book “What If” by Randall Munroe has a chapter called No More DNA dedicated to what would happen if someone had all of their genes instantly disappear. I imagine a toxin or weapon that knocks down genes would have a similar effect.
Transcription control
The fact that the Cas9-crRNA-tracrRNA can bind to a specific site can be exploited for transcription control, as well. Using Cas9 protein mutants that are engineered to lose their endonuclease activity will prevent it from cutting the DNA. However, the protein will still bind to the site complementary to the crRNA target sequence and then…it will just sit there. But this can block the RNA polymerase needed to transcribe, and thereby express, the gene from doing its job. The Cas9-crRNA-tracrRNA complex binding can then be reversed, removing it from the target site, thereby allowing the gene to be expressed. This reversible association can be exploited in research to study the function of genes without having to actually permanently alter the genome.
Science fiction idea: some sort of injection that, say, a soldier could use that would activate and/or repress certain genes to give them temporary affects. Think Berserkers from ancient times by temporarily increasing adrenaline and metabolism. Could also possibly be used as a drug by temporarily increasing dopamine production.
Germline genetic engineering
The idea of designer babies is not only a science fiction trope in movies like Gattaca anymore. Researchers in Guangzhou, China and in Oregon, U.S.A. have already used CRISPR/Cas9 to edit the human genome in embryos. This could lead to things like the eradication of childhood diseases like Tay-Sachs, Cystic Fibrosis, Huntington Disease, and so many more.
Germline genetic engineering could also, theoretically, lead to the ability to give certain traits to one’s offspring by genetically modifying an embryo before implantation in the womb. Things like hair color, eye color, skin color, height, strength, intelligence, creativity, interests, and even morality could, theoretically, be manipulated.
Obviously, this leads to both practical and ethical issues. For instance, as far as practical issues go, what if evolution has settled on our current range of intelligence for a reason? What if having significantly better working memory somehow clutters the mind too much to function properly?
Somatic cell genetic modification
Being able to edit the genome of an embryo is all fine and good, but that only helps my potential offspring. What about me? Is it possible to edit my own genome? Once again, this sounds like a science fiction idea, but gene therapy has been around since the 1990s. Viruses have been used for gene therapy because a virus can be very specific for a certain type of cell, but it is difficult to control where the genetic material ends up in the genome. The CRISPR/Cas9 system could be the answer.

There are, unfortunately, various issues with this. In a laboratory, using CRISPR/Cas9 to edit the genome of cells is a crap shoot. Millions of cells are treated, but only a few will actually be successful, and various selections methods have to be used in order to isolate those cells and use them to start a cell line (getting many more cells descended from the successfully treated cell such that they all have the same modification). I have not yet personally used the CRISPR/Cas9 system, but others in my lab have, and it can be a long, frustrating process trying to get it to work. Granted, the techniques are still in their infancy (as of 2018 it has only been five years since the Cong et al 2013 paper demonstrating the proof of concept use of CRISPR/Cas9 for targeted gene editing). Who knows what the next 5, 10, 20, 50, 100+ years of scientific progress will bring?
Gene Drive
This is a method that could be utilized for things like eliminating mosquitoes that carry malaria or zika or ticks that carry Lyme disease. I don’t think I could do a better job of describing this method than the following video.
Conclusion
The CRISPR/Cas9 system is an amazing recent discovery that not only has the potential to revolutionize biotechnology as we know it, but it is also a deep well of plausible ideas for science fiction writers with a keen eye for accuracy and thoroughness. I hope that this blog post can be helpful for anyone, regardless if you are a sci-fi writer, reader, or just interested layperson.
—
Post Script

The featured image of Jessica Jones was taken from this blog. I have to point out, though, that the DNA strand shown in the image has a left-hand helical turn (do a thumbs up on your left hand: as you move along the DNA strand, the helix spirals in the direction your fingers are pointing). In reality, DNA has a right-hand helical turn.

This seems nit-picky, but it is also very important biochemically, and a lot of movies and TV shows use the left-handed and right-handed helices interchangeably. Ever since I learned about this as an undergrad, I haven’t been able to un-see it. Now you won’t be able to, either.
